Biological infinitezimal - organic databases
THE BIOLOGICAL INFINITE - ORGANIC DATABASES
Dr. chem. eng. Ciprian-Gabriel CHISEGA-NEGRILĂ
Motto:
In the beginning it was simplicity. It is quite difficult to explain the beginning of the universe, however simple it may have been. I assume everyone accepts the idea that it is even more difficult to explain the sudden appearance, with all the necessary equipment, of the complex order - life, or of a being capable of creating life. Darwin's theory of evolution by natural selection is satisfying because it shows us a way in which simplicity can turn into complexity, how some disordered atoms can combine into increasingly complex structures, until they end up creating humans.
Richard Dawkins – The Selfish Gene, Chapter 2 – The Replicators, pg. 14, Bucharest, Technical Ed., 2001 (trans. Dan Crăciun)
I. ABSTRACT
RO: The article mainly describes the structure of the eukaryotic cell along with sub-structures in the cytoplasm and nucleus to facilitate understanding of how deoxyribonucleic acid/DNA functions as a database containing all the information the cell needs to thrive and multiply. It also describes the mechanism by which the information contained in DNA is transcribed into ribonucleic acid/RNA and then used as templates for protein synthesis within the ribosome. The architecture of the article is one of metatext describing fundamental level information about the eukaryotic cell – the building block of higher organisms (which includes humans). For a better understanding of the treated subject, QR-codes representing links to in-depth information and bibliographic references have been inserted into the text.
ENG: The article primarily describes the structure of the eukaryotic cell along with the sub-structures within the cytoplasm and nucleus to facilitate understanding of how deoxyribonucleic acid (DNA) functions as a database containing all the necessary information for the cell to thrive and reproduce. It also explains the mechanism by which the information contained in DNA is transcribed into ribonucleic acid (RNA) and then used as a template for protein synthesis within the ribosome. The architecture of the article is one of metatext, providing fundamental information about the eukaryotic cell—the building block of higher organisms (including humans). For a better understanding of the subject, QR codes have been included in the text, leading to in-depth information and bibliographic references.
Keywords: living organism, cell, DNA, RNA, protein, database
II. INTRODUCTION
The planet we all live on has a history of about 4.5 billion years behind it. Will it be much? Will it be a little? The fact is that it is almost a third of the age of the Universe. The first forms of life appeared about 4 billion years ago in the form of single-celled prokaryotic organisms. This cell type includes bacteria and members of the class Archaea. They are distinguished by the fact that they do not have a nucleus, but they have a flagellum (most of the time), a cell membrane, cytoplasm, ribosomes and nucleotides that contain the genetic information dispersed in the cytoplasm.
II.a. EUKARYOTIC ORGANISMS
Eukaryotic organisms are much more complex organisms than prokaryotes. Compared to prokaryotes, eukaryotic cells are about 10,000 times larger. Both unicellular and multicellular eukaryotes can be included in this category. They are cells that belong to animals (such as humans), plants (such as corn) and fungi (such as Champignon mushrooms).
Other representatives of this class are: molds, protozoa and algae.
II.b. COMPONENT PARTS OF THE EUKARYOTIC CELL
The eukaryotic cell is described in Fig. 1.
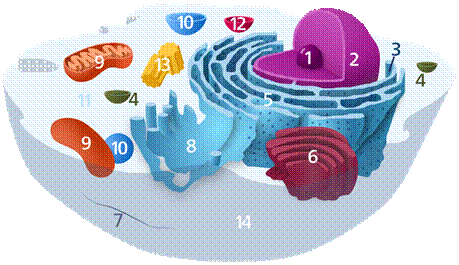
Fig.1 – The structure of a eukaryotic cell: 1 – nucleolus, 2 – nucleus, 3 – ribosome, 4 – vesicle, 5 and 8 – endoplasmic reticulum, 6 – Golgi apparatus, 7 – cytoskeleton, 9 – mitochondrion, 10 – vacuoles, 11 – cytosol, 12 – lysosome, 13 – centrosome, 14 – cell membrane
It is composed of the following components (Fig.1):
- Membrane (14) – it is the one that covers the cell, controls what enters or leaves it and maintains its electrical potential;
- Cytoplasm – is a gel-like environment that occupies most of the space bounded by the membrane. It contains the intra-cytoplasmic sub-structures described in the following 12 items:
- The nucleolus (1) is the largest structure in the nucleus of eukaryotic cells. It is known to be the site where ribosome biogenesis takes place;
- Nucleus (2) – contains almost 99% of the cell's genome/database. This is where deoxyribonucleic acid – nuclear DNA is found (absent in red blood cells/erythrocytes). It is organized/packaged mostly in the form of chromosomes. All the information needed by the cell is stored here. We will return to this topic further!
- Ribosome (3) – a sub-structure of the cytoplasm where protein synthesis takes place using DNA/RNA information as a template;
- Vesicles (4) – structures inside or outside a cell that contain a fluid or cytoplasm enveloped by a double layer of lipids. It helps in the removal or inclusion of substances from/into the cell, as well as the transport of substances across the membrane;
- Endoplasmic reticulum (5 & 8) – is part of the transport system of the eukaryotic cell and has, among other things, the function of protein folding/packaging;
- The Golgi apparatus (6) – has a very important role in embedding proteins in vesicles before they are sent to their destination;
- Cytoskeleton (7) – is a complex and dynamic network of interconnected protein filaments. Inside eukaryotic cells it extends from the nucleus to the cell membrane. Gives the cell resistance to deformation. It is also involved in many other cellular processes.
- Mitochondria (9) – are used to generate and store energy in chemical form in ATP (adenosine triphosphate) molecules;
- Vacuoles (10) – are closed compartments in the cytoplasm and can contain inorganic or organic substances (often enzyme solutions, and in rare cases solid substances);
- Cytosol (11) – also known as the cytoplasmic matrix. It is a complex mixture of substances dissolved in a large amount of water;
- Lysosomes (12) – They are spherical vesicles that occur in many animal cells. They contain hydrolytic enzymes that digest all kinds of biomolecules;
- Centrosomes (13) – is an organelle with a complex function.
III. DESOXYRIBONICLEIC ACID – DNA – ORGANIC DATA BASE
DNA is a very complex organic molecule, but I will try to explain it as simply as possible! It was discovered (he, DNA ... ☺) at the beginning of the 19th century by the Swiss biochemist Frederich Miesher, but its structure was deciphered only in 1953 by James Watson, Francis Crick, Maurice Wilkins and Rosalind Franklin who studied this molecule with the help of a new technique for that time - X-ray diffraction. The four scientists showed that DNA consists of two twin helices "twisted" around each other, so the so-called α-helix.
The place where DNA resides is in the cell nucleus (Fig.1) where it is packaged (and super-archived) in the form of several X-shaped structures that represent chromosomes. Specifically, in humans, there are 46 such chromosomes, grouped into 23 pairs. Twenty-two such pairs are called autosomes and have a similar structure in both females (♀) and males (♂), and the 23rd pair differs because it encodes information about the organism's sex.
All this packing that I was talking about before means that a molecule that, left free, has a "huge" length of almost 2 meters (molecules usually have dimensions of the order of nano- or even micrometers) can be archived in a space of the order of nanometers (this is about the size of the nucleus of a normal eukaryotic cell). Packaging is done with the help of small molecules called histones. For clarity of explanation I provide a detailed graphic representation in Fig.2.
It is observed that on the skeleton made of sugar molecules (deoxyribose, for completeness ... a sugar similar to ribose in RNA ... but which has lost a hydroxyl -OH group) and phosphate-, Fig.3-4 (derived from ATP – the adenosine triphosphate we were talking about earlier when the mitochondrion was briefly presented: position (9) of Fig.1) are the nuclear bases, the 4 letters of the genetic alphabet: A – adenine, C – cytosine, G – guanine and T – Thymine (Fig.4). These characters from the genetic alphabet combine two at a time to link the two strands of the DNA molecule. Adenine (A) with Thymine (T), through 2 hydrogen bonds, A === T and Cytosine (C) with Guanine, through 3 hydrogen bonds, C ≡≡≡ G (Fig.3-4).
Several nuclear bases in a sequence make up a gene, and genes are what contain all the information a cell needs to thrive and multiply. DNA replication when the cell reproduces is not the subject of this article...
Many of these genes code for proteins (long sequences of amino acids strung one after the other) which can be: enzymes, transport substances, hormones or even neurotransmitters (as for example in the synaptic space between 2 neurons). DNA also encodes information necessary for the growth and development of an egg fertilized by a spermatozoon until the moment of obtaining a fully functional organism … But about that in the next chapter …
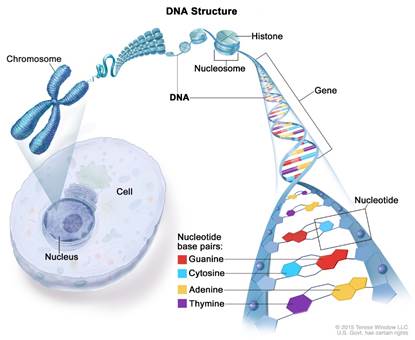
Fig.2 – Location of DNA in the nucleus and its packaging
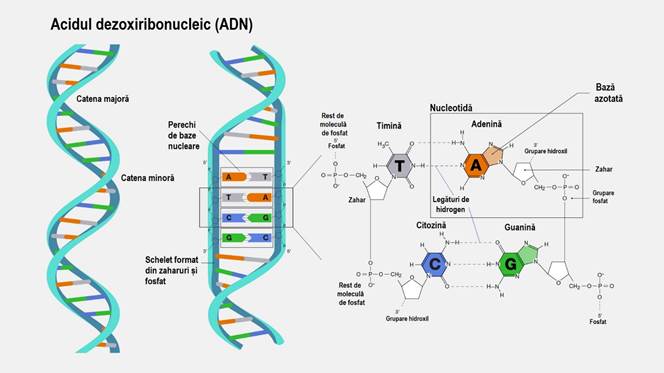
Fig.3 – Detail that includes the two coiled helices (α-helix) and the nuclear bases: adenine (A), thymine (T) which binds through 2 hydrogen bonds A === T, cytosine (C), guanine (G) which binds through 3 hydrogen bonds C ≡≡≡ G
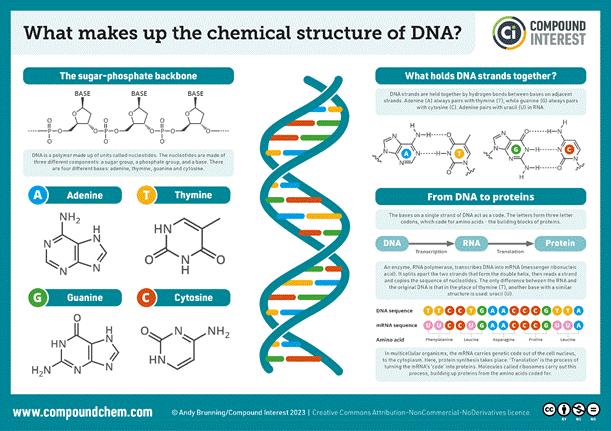
Fig.4 – The structure of the nuclear bases and the skeleton of the two main chains
IV. RNA – RIBONUCLEIC ACID INTRACELLULAR TRANSPORTER
RNA is a slightly simpler molecule than DNA. If DNA has two strands wrapped around each other, RNA has only one strand. The sugar in the backbone of the RNA strand is called ribose, as described in the previous subsection. The RNA alphabet still has 4 characters, but one base is different from those in DNA. Thus, there are: adenine (A), cytosine (C), guanine (G) and uracil (U). Note that thymine no longer appears in this alphabet. As such, when transcribing DNA into RNA, thymine is replaced by uracil: adenine is transcribed as uracil, cytosine as guanine, guanine as cytosine, and thymine as adenine.
The main role of RNA is to carry information from the DNA in the nucleus to the ribosome in the cytoplasm of the cell, where it is used to synthesize the proteins that the cell needs at any given time.
There are several types of RNA:
- Messenger RNA – carries the coding information from DNA to the ribosome;
- Transfer RNA – is involved in the mechanism by which amino acids are linked in an orderly manner into a protein (inside the ribosome) … but I won't go into details here;
- ARN ribozomal, etc.
For the transposition mechanism of the information from the coding genes on the DNA (in eukaryotes - including humans - almost 97% of the DNA information has no role in protein synthesis) I will continue to discuss only messenger RNA - mRNA.
V. TRANSCRIPTION OF INFORMATION FROM DNA INTO PROTEINS (amino acid sequences)
As we said before, messenger RNA - mRNA - is the carrier of information inside the cell. When it is necessary to transcribe information from a gene, an enzyme called RNA polymerase is attached to the DNA structure. This unwinds the two strands and leaves the nuclear bases 10 – 20 characters apart (A,T, G, C). The chain from which the transcription is made is called the template (antisense) chain, and the other nontemplate (coding). The bases on DNA are transcribed as before: adenine (A) as uracil (U), cytosine (C) as guanine (G), guanine (G) as cytosine (C) and thymine (T) as adenine (A).
For example (and here we took 27 characters, but the genes that are transcribed are usually much longer):
If on the coding chain we have:
| A | T | G | C | G | G | C | A | C | G | A | T | T | T | C | C | T | G | A | A | C | C | C | G | T | G | A |
on the correspondence template chain produces:
| T | A | C | G | C | C | G | T | G | C | T | A | A | A | G | G | A | C | T | T | G | G | G | C | A | C | T |
and the RNA strand is transcribed:
| A | U | G | C | G | G | C | A | C | G | A | U | U | U | C | C | U | G | A | A | C | C | C | G | U | G | A |
which is called mRNA or messenger RNA.
At this point, we move on to mRNA processing because not all information is coding. Some parts of the mRNA will be excluded and these are INTRONS and the rest of the sequence that remains are called EXONS. These exons come together and form the information that will reach the ribosome for protein synthesis. Here the "letters" on the sequence are read 3 at a time and expressed as a single amino acid. There are 64 combinations of these characters and only 20 amino acids, which means that an amino acid can be coded by several groups of 3 characters. There is a START combination: AUG and 3 STOP combinations: UAA, UAG and UGA. But amino acids will be discussed in the next sequence of characters in the Latin alphabet!
V.a. AMINO
There are 20 amino acids in human genetics, each with its own chemical formula, as described in Fig.5 and in Tab.1 from the sequel. What all these amino acids have in common: basic amino groups -NH2 and carboxylic acid groups -COOH (where R represents a certain radical not specified here).
Due to this peculiarity of the amino acid molecule, several such molecules can bind to each other to form long chains of the type:
H2N – C(R1) – (C=O) – HN – C(R2) – (C=O) – HN – C(R3) …….. (C=O) – HN – C(Rn) – COOH …. and ... look, this is how I "bewitched" you until I described how proteins are formed ☺! In the biological or rather biochemical system, things are a little more complicated than that, but I will explain in more detail in the following paragraphs.
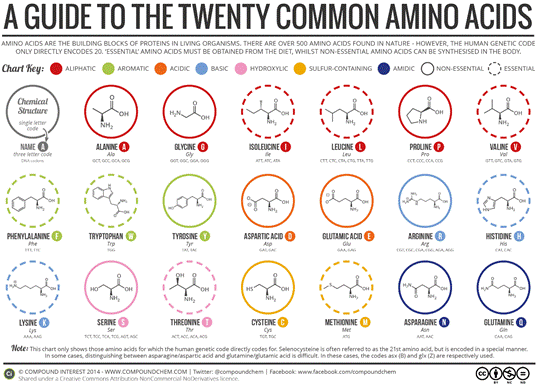
Fig.5 – The most common amino acids
Tab.1 – Amino acid names, symbols and electrical charge of the molecule
| Name | 3 letter symbol | Symbol 1 letter | Polarity | Coding combinations on mRNA |
| Alanine | Ala | A | Neutral | GCU, GCC, GCA, GCG |
| Arginine | Arg | R | (+) | CGU, CGC, CGA, CGG, AGA, AGG |
| Asparagine | Asn | N | Neutral | AAU, AAC |
| Acid aspartic | Asp | D | (-) | FALSE, GAC |
| Cysteine | Cys | C | Neutral | UGU, UGC |
| Acid glutamic | Glu | E | (-) | GAA, GAG |
| Glutamine | Gln | Q | Neutral | CAA, CAG, GAA, GAC |
| glycine | Gly | G | Neutral | GGU, GGC, GGA, GGG |
| Histidine | His | H | 10%+, 90%- | CAU, CAC |
| Isoleucine | With | I | Neutral | OH, OH, OH |
| Leucine | Leu | L | Neutral | UUA, UUG, CUU, CUC, CUA, CUG |
| Lysine | Lys | K | (+) | AAA, AAG |
| Methionine | Met | M | Neutral | AUG |
| Phenylalanine | Phe | F | Neutral | UUU, UUC |
| Proline | Pro | P | Neutral | CCU, CCC, CCA, CCG |
| Serene | To be | S | Neutral | UCU, UCC, UCA, UCG, AGU, AGC |
| Threonine | Thr | T | Neutral | ACU, ACC, ACA, ACG |
| Tryptophan | Trp | W | Neutral | UGG |
| Tyrosine | Tyr | Y | Neutral | UAU, UAC |
| valine | Val | V | Neutral | GUU, GUC, GUA, GUG |
| START | AUG | |||
| STOP | UAA, UAG, UGA |
If on the mRNA chain we have (the example from before):
| A | U | G | C | G | G | C | A | C | G | A | U | U | U | C | C | U | G | A | A | C | C | C | G | U | G | A |
in the language of amino acids this translates as (a combination of 3 characters codes for an amino acid):
| START | Arg | His | Asp | Phe | Leu | Asn | Pro | STOP |
| START | R | H | D | F | L | N | P | STOP |
that is:
START – Arginine – Histidine – Aspartic acid – Phenylalanine – Leucine – Asparagine – Proline – STOP
or as polarities:
| START | (+) | 10%(+)90%(-) | (-) | neutral | neutral | neutral | neutral | STOP |
The information in Tab.1 is summarized graphically in Fig.6 from the following:
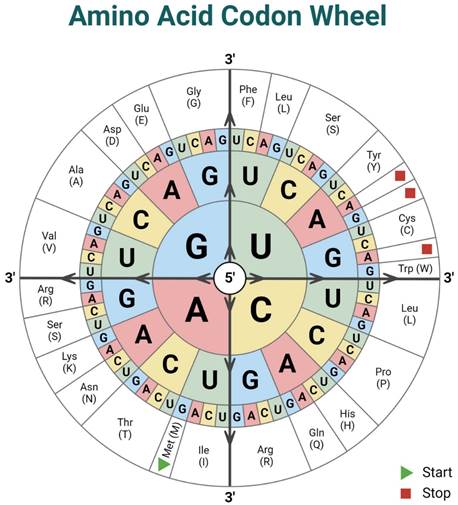
Fig.6 – The codon wheel
Why have we insisted so much on DNA, RNA, mRNA, amino acids or amino acid sequences (more broadly called proteins)? Because, these amino acid sequences have a function, which function is given by the way they fold or arrange themselves spatially. And this 3D arrangement is done according to the polarity of each amino acid present in that protein. Because the environment in the cell cytoplasm is aqueous (therefore polar), the 3D structure appears following several criteria:
- a non-polar amino acid will want to hide as deep as possible inside the structure because it doesn't like water;
- a polar amino acid will want to sit as close as possible to one with the opposite polarity;
- a polar amino acid will want to stay as far as possible from an amino acid of similar polarity.
And this spatial conformation is what dictates the appearance of an - active site - that allows coupling to another protein, to an enzyme (which makes a process run faster and with less energy consumption), to a molecule of active substance in a medicine. Because drugs (most of the time) work on the principle of a false key that fits to open a specific lock/active site of a protein.
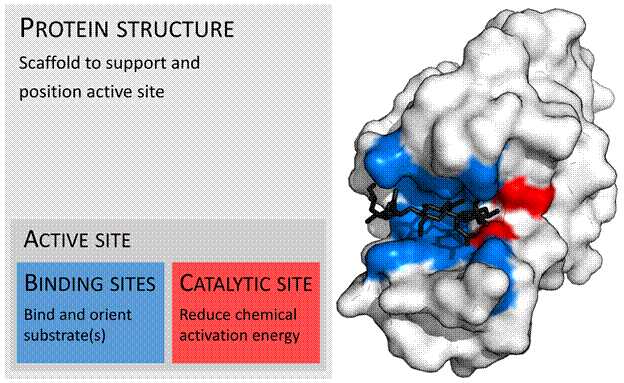
Fig.7 – Active site of a protein that appeared as a result of its arrangement in 3D
For those who want to delve deeper into the subject, I have left the "candy on the cake" for the end, some explanatory videos.
VI. Instead of CONCLUSIONS – A HYPOTHESIS!
At one point, I was saying that: DNA encodes information necessary for the growth and development of an egg fertilized by sperm until the moment of obtaining a fully functional organism! This is a certain and certain thing as can be seen in Fig.8.
From an article written by Manuela Murariu and Gabi Drochioiu and published in the journal Biosystems (IF 2.0) in 2012, I learned about the biostructural theory in living organisms. They talk about the supramolecular conception in biology and how Eugen Macovschi tried, and partially succeeded, to explain the differences between "dead" and "alive" in biological systems.
The hypothesis I want to launch is:
Information that is not related to protein synthesis (in humans – almost 97% of DNA information has no role in protein synthesis, but also encodes information for the growth and development of an embryo to the stage of a fully functional organism, Fig.8) DO THOSE BIOLOGICAL SUPERSTRUCTURES ALSO CODE WHAT MAKES THE DIFFERENCE BETWEEN LIVING AND NON-LIVING? DO THESE CODE BIOFIELDS?
Here to see you!
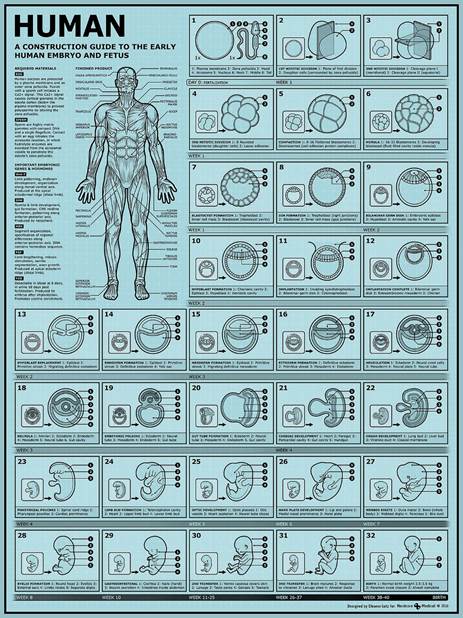
Fig.8 – The development of the human embryo from the moment of conception until birth
VII. SELECTIVE BIBLIOGRAPHY
For the ease of citing, but also of tracking information in the online environment, we have chosen that web pages that provide additional information are cited inside the text in the form of QR codes.
Dawkins, R. – Gena Egoistă, Bucharest, Technical Ed., 2001 (trans. Dan Crăciun)
Darymple, G.B., – The Age of th Earth, Stanford, Stanford University Press, 1991;
Manhesa, G., Allègre, C.J., Dupréa, B., Hamelin, B. – Lead isotope study of basic-ultrabasic layered complexes: Speculations about the age of the earth and primitive mantle characteristics – Earth and Planetary Science Letters, 1980;
Dinu, V., Trutia, E., Popa-Cristea, E., Popescu, A. – Medical Biochemistry, Small treatise, Bucharest, Medical Ed., 1996
Murariu, M., Drochioiu, G. – Biostructural theory of the living systems – Biosystems, 109-2, 126-132, 2012


